I tried to copy EIC Signal description items near each other and I don't know how to avoid this problem showed below.
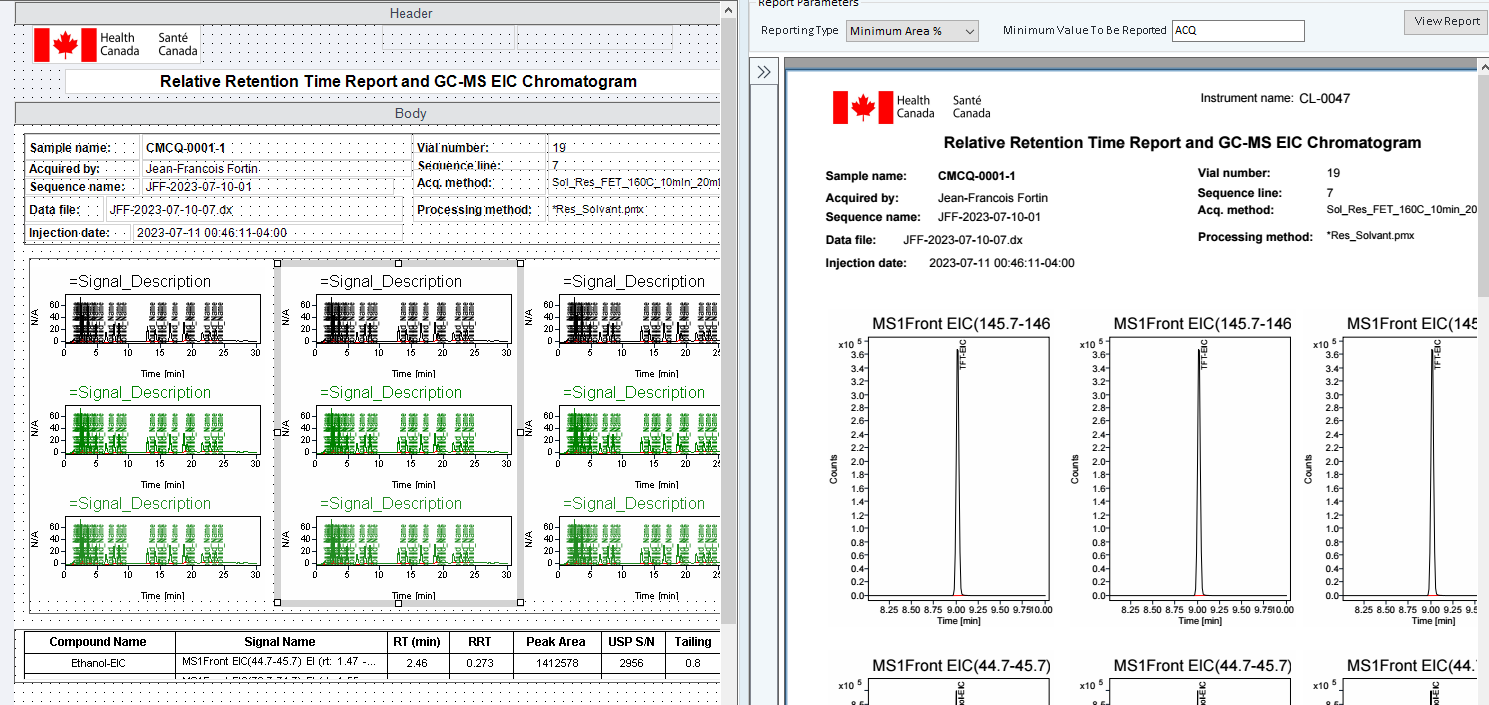
As you can see, the three chromatogram is the same compound. How can I avoid this repetition?
thank you for your help.
I tried to copy EIC Signal description items near each other and I don't know how to avoid this problem showed below.

As you can see, the three chromatogram is the same compound. How can I avoid this repetition?
thank you for your help.
Hello,
Try the MS Chromatogram Flowlayout object instead of the individual chromatograms.
Marty
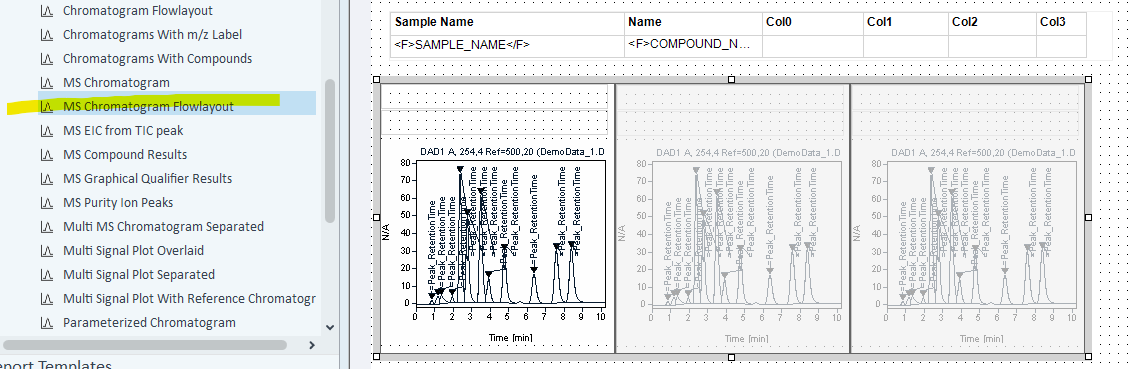
Hi Martin. I have the same problem. I don't know how to manage the group repeating so we can have different compound name on chromatogram lighted in gray.
Ok I tried to put my filter on the outside group. It's getting better but not completely OK.
Here is the outside filter performed with a sorting based on Peak retention time :
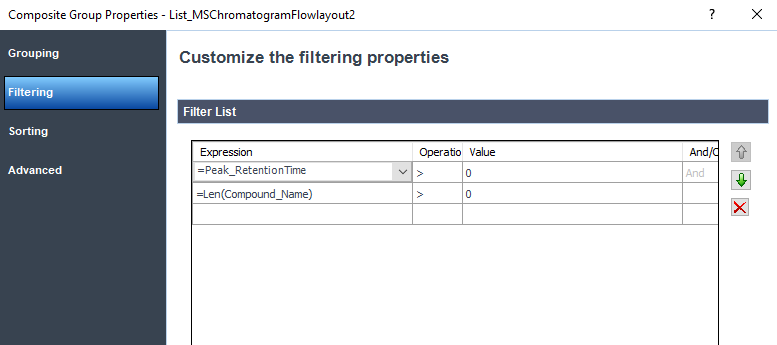
I erase the Len(compound_name) filter at the chromatogram and there is no sorting for this section.
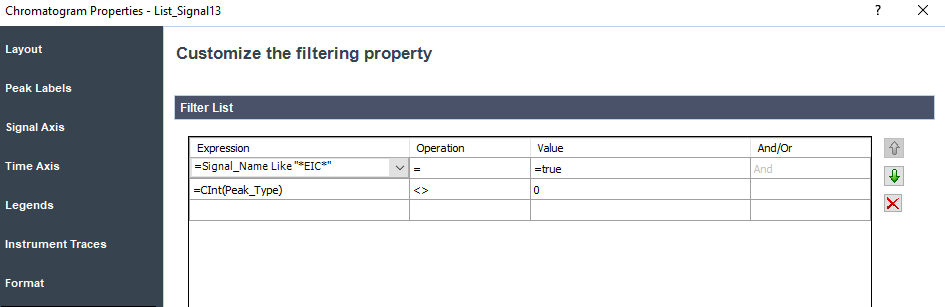
Here is the results:
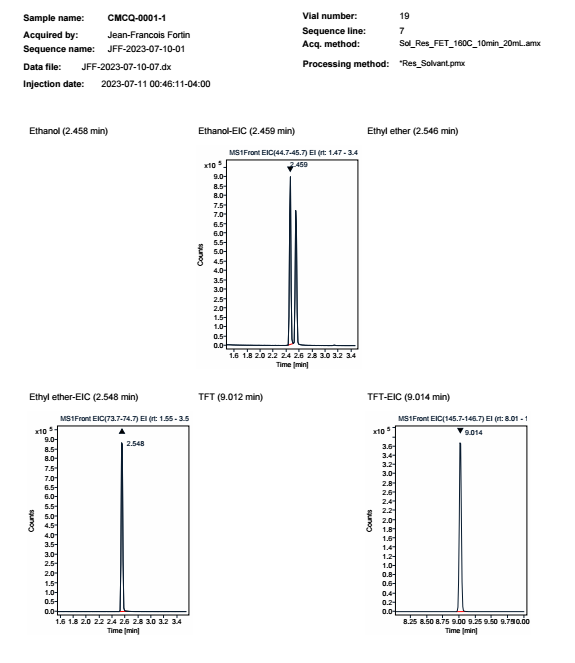
We can see that the filter EIC on the chromatogram doesn't work as final result as we can see blank chromato with no EIC as name compound.
I will try to add the EIC filter again on the outside filter.
it works! Even if I erase the EIC filter at the chromatogram portion. Thank you for your help. I will saved this composite group. If I add this composite group into a sequence summary report instead of single injection report, would you recommend to ass sorting based on injection ID?
I finnaly got what I have wanted. Thank you again. I tried to add the Multiplier used in the calibration Compound table used for a specific calculation with samples into my sequence injection summary:
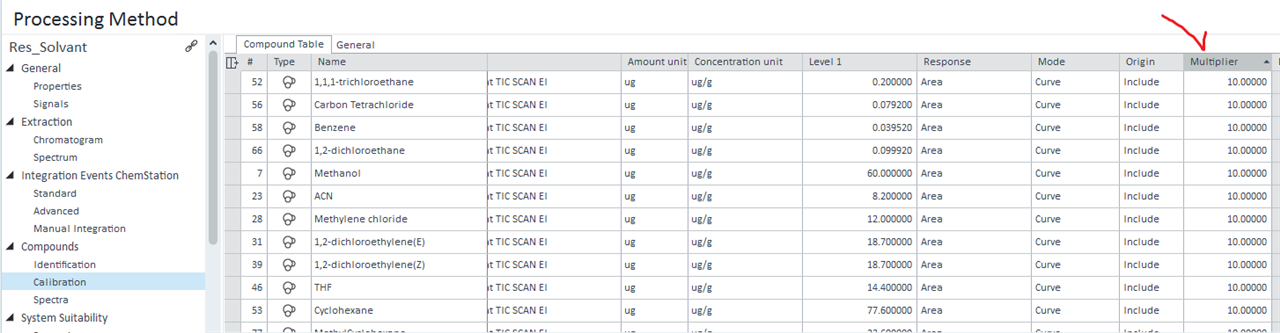
But I only found multipliers that included the 5 possible multipliers.
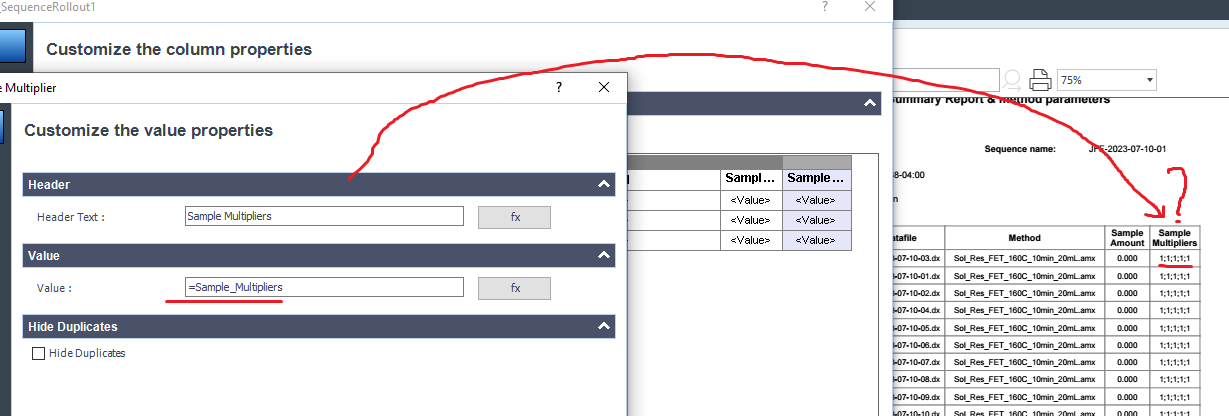
Do you know how to select a specific multiplier to add on the sequence summary?
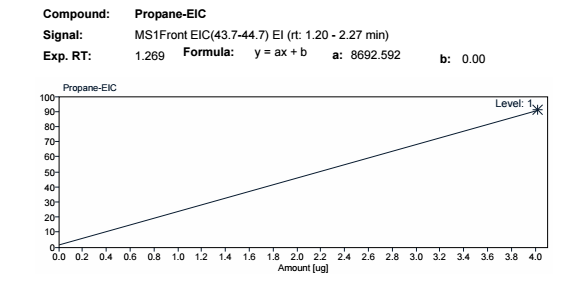
Do you know how to change the response value of the courve into raw date ms counts?
Right now it's the 100% of response relatec to the 1 calibration point.
Hello,
The explanation from the Openlab Help and Learning is copied below. Depending on where you are using the resultant value you may need to place the expression in a Val() function.
Marty
Depending on the value of a data field, you may want to show only part of the value.
For example, the data fields Sample_DilutionFactor and Sample_Multipliers show five numbers separated by semicolon, but only part of the numbers may be set by the chromatographic data system. Therefore the value may look like 10; 0; 0; 0; 0 if only the first number is set. To display only the number 10 instead of the entire string, you can use the following expression:
=Choose(1, Split(Sample_Multipliers, ";"))
The Split function divides the string in several parts, using the semicolon as a delimiter. In this example, the different parts are the single numbers.
The Choose function selects and returns a specific value from a list of values. In this example, it returns the first value, that is, the number 10.
it works but I obtained a value of 1. It is supposed to be 10. I put 10 as multiplier in the calibration table and it also works to calculate concentration in ppm (ug/g).
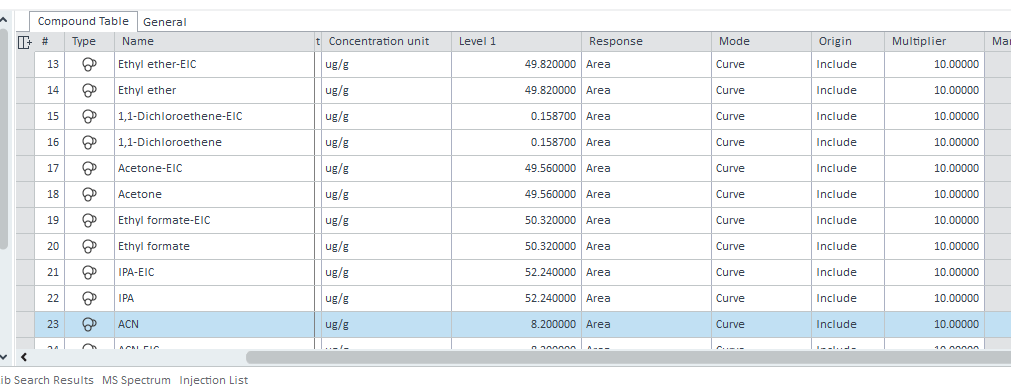
But in the injection list, the five multiplier are at 1.
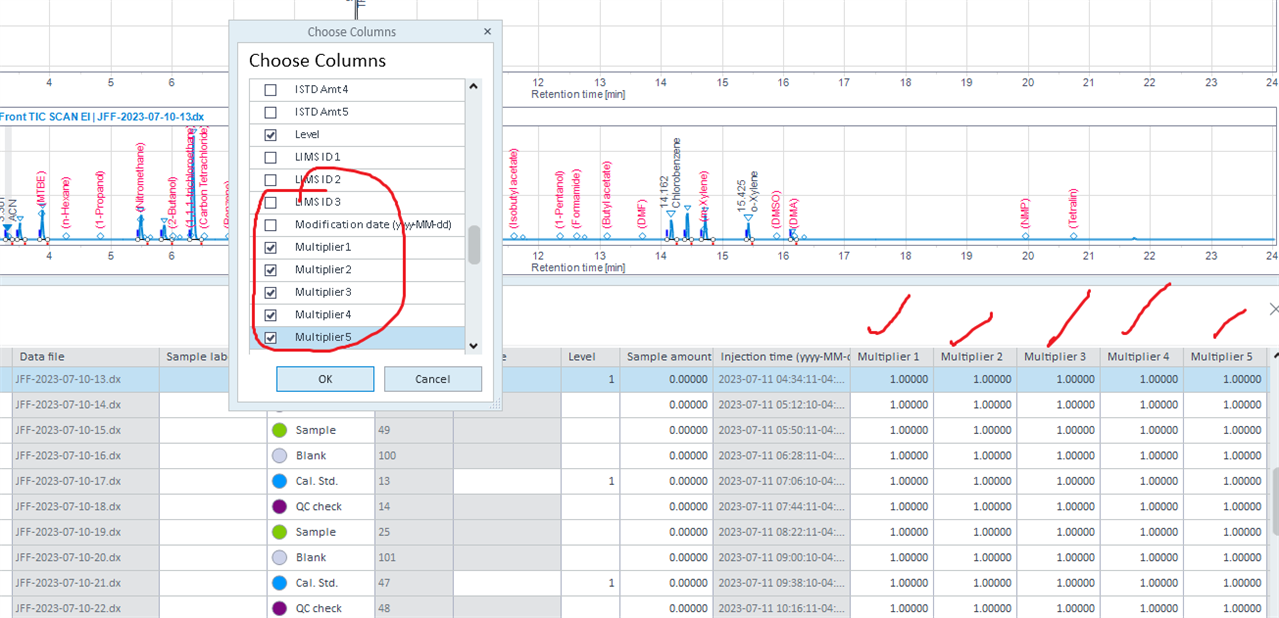
I am not sure what's wrong here.
I need to use the function related to the multiplier used in the process method.
Which multiplier is used in the concentration mass % ?
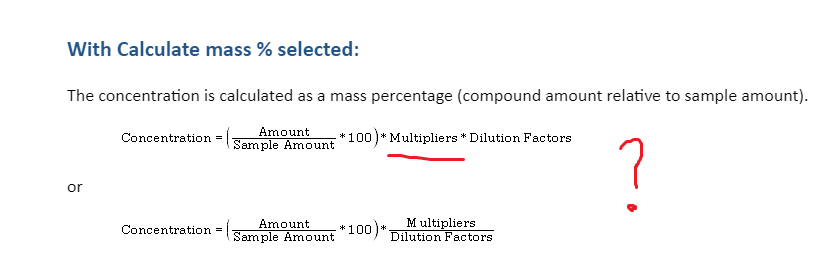

Based on my results, the multiplier need to be a 10 in order to give my result in ppm. It seems to work. I will do a test tomorrow addind another multiplier and check if my sample's results are affected.
it works but I obtained a value of 1. It is supposed to be 10. I put 10 as multiplier in the calibration table and it also works to calculate concentration in ppm (ug/g).

But in the injection list, the five multiplier are at 1.

I am not sure what's wrong here.
I need to use the function related to the multiplier used in the process method.
Which multiplier is used in the concentration mass % ?

Based on my results, the multiplier need to be a 10 in order to give my result in ppm. It seems to work. I will do a test tomorrow addind another multiplier and check if my sample's results are affected.
Hello,
All the multipliers are used in the concentration calculation. The multiplier in the method is the compound multiplier which is different from the sample multipliers 1-5. The compound multipliers can be different for each compound in the peak table but are the same for all injections. The sample multipliers are applied to all results for a given injection but can be different for each injection
Marty.
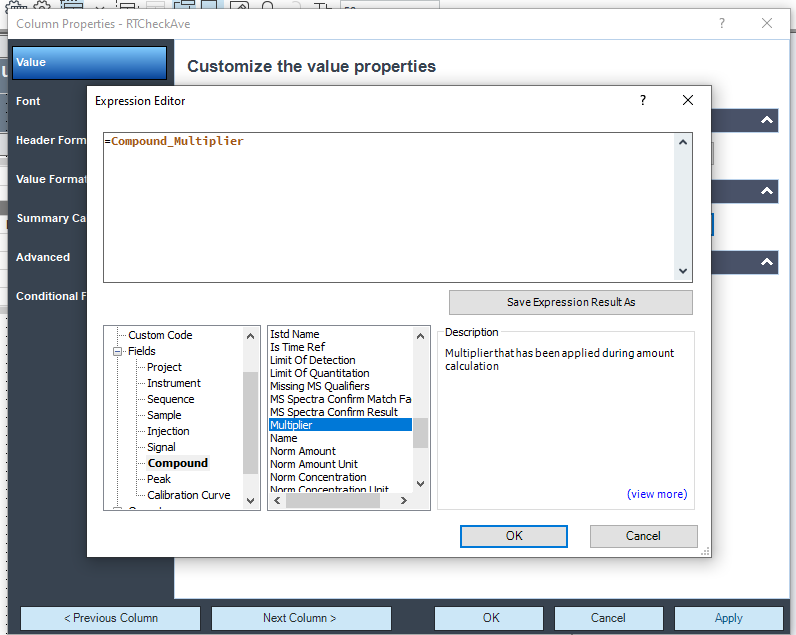
It's weird because I don't have this option on my side. No Multiplier.
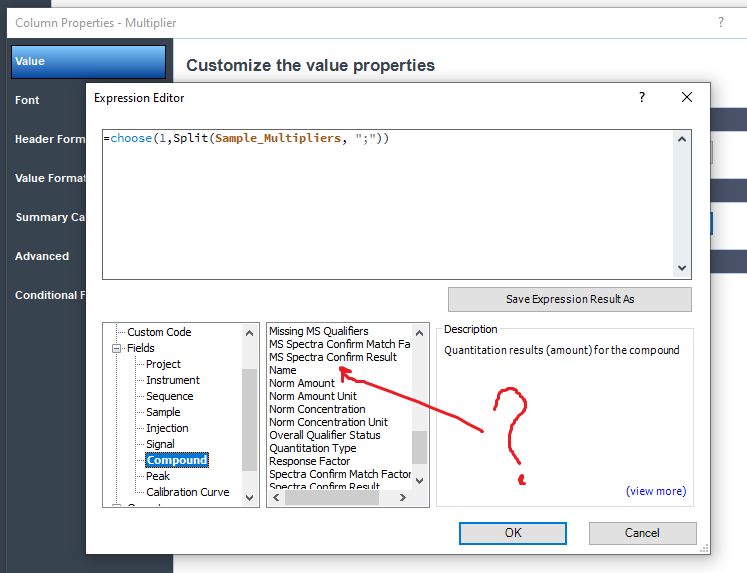
In my version, that field isn't available. However, he can get it by the calculation =Compound_Concentration/Compound_Amount
I don't have the multiplier field either on my openlab CDS 2.6.
I don't understand your calculation to retrieve my multiplier of 10.
if I use your equation with my first example it will be :
393.60 ug/g divided 9.84 = 40. It doesn't work. it's supposed to be 10.
Does it return exactly 40? If you pump up the number of decimals is it 40.000000000? If so, then it's probably using one or multiple values you entered somewhere, which would allow you to modify the calculation to account for that. At least on my end it just spits out the multiplier, but I also doubt I have all the same stuff filled out as you do.
Did you enter a 4 somewhere? Or a 0.25?
My equation is :
ug solvant/g samples= amount/sample weight (mg) * 100 *10
In bold view, this is calculated with the % mass formula. the multiplier 10 is the one I want to show on my sequence table summary.