I tried to copy EIC Signal description items near each other and I don't know how to avoid this problem showed below.
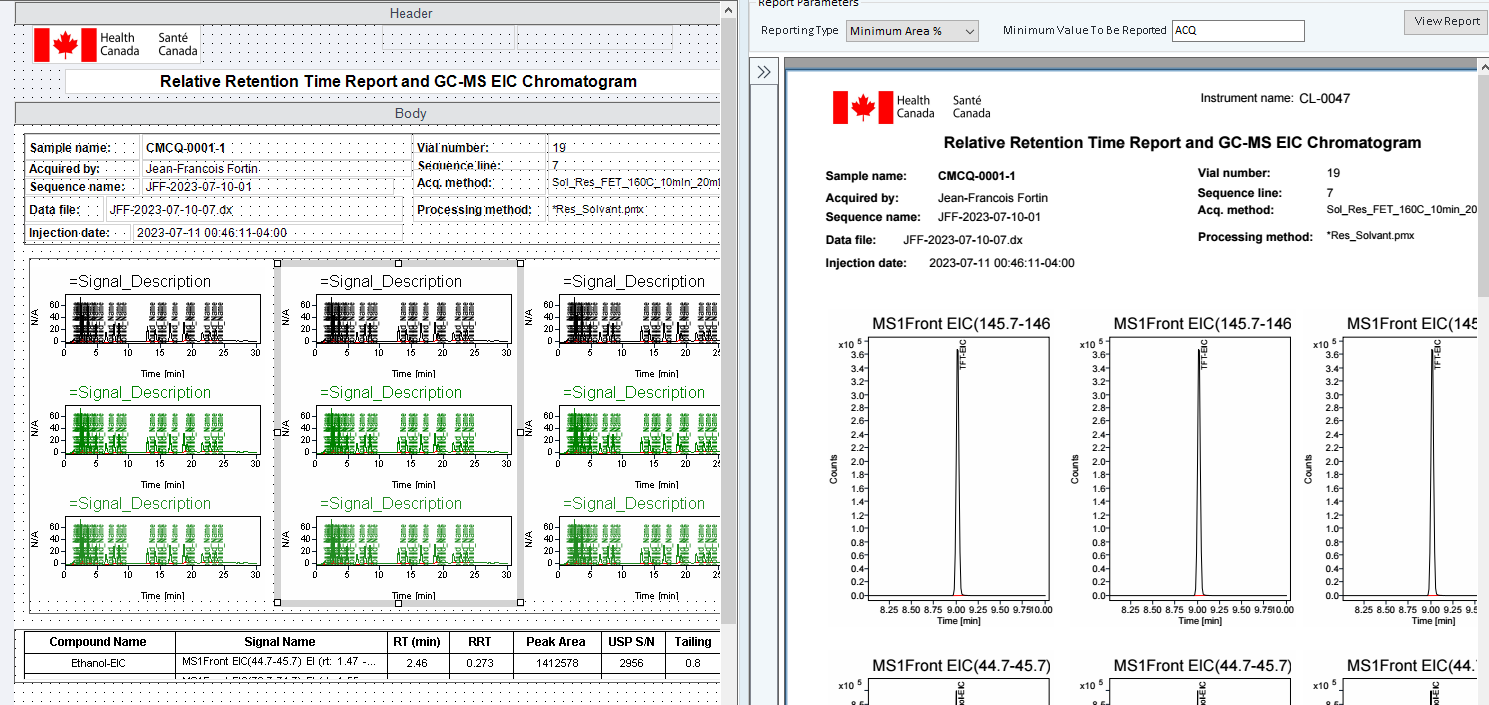
As you can see, the three chromatogram is the same compound. How can I avoid this repetition?
thank you for your help.
I tried to copy EIC Signal description items near each other and I don't know how to avoid this problem showed below.

As you can see, the three chromatogram is the same compound. How can I avoid this repetition?
thank you for your help.
Hello,
Try the MS Chromatogram Flowlayout object instead of the individual chromatograms.
Marty
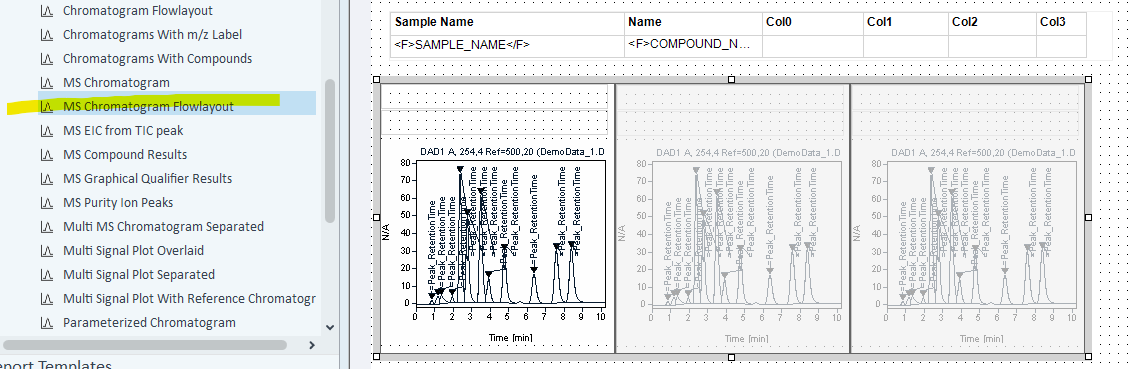
Hi Martin. I have the same problem. I don't know how to manage the group repeating so we can have different compound name on chromatogram lighted in gray.
Hello,
It looks like your filter is on the chromatogram object not the outside group. See the example below where I added Len(compound_name) > 0 as a filter in both locations.
Filter on the Chromatogram object
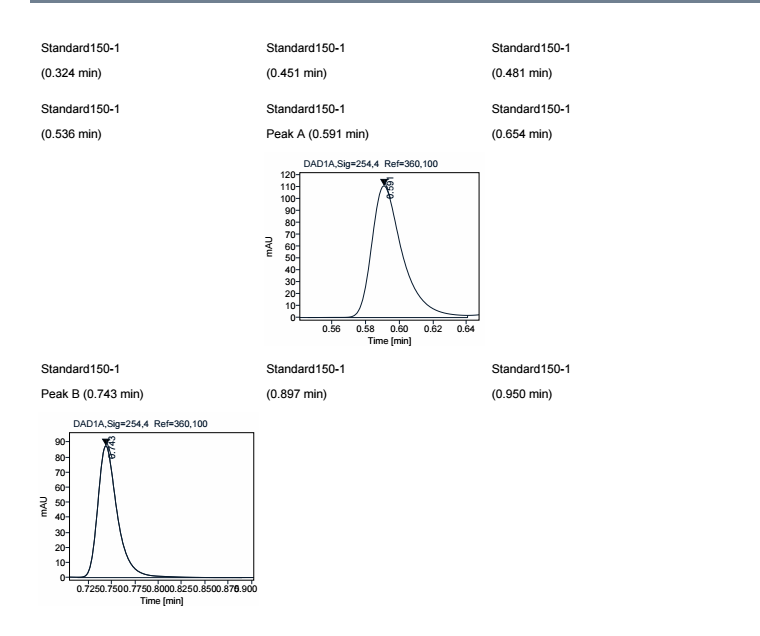
Filter on the group object
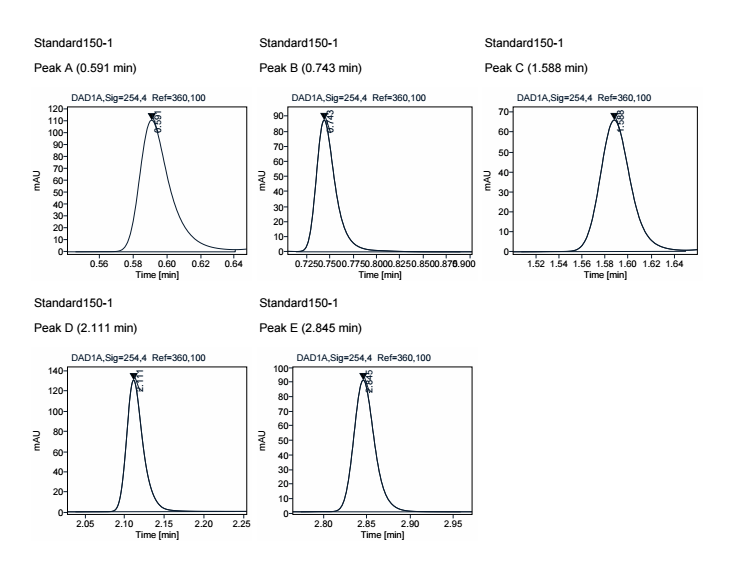
Oh Perfect, I will try this. Thank you again for your precious help.
Ok I tried to put my filter on the outside group. It's getting better but not completely OK.
Here is the outside filter performed with a sorting based on Peak retention time :
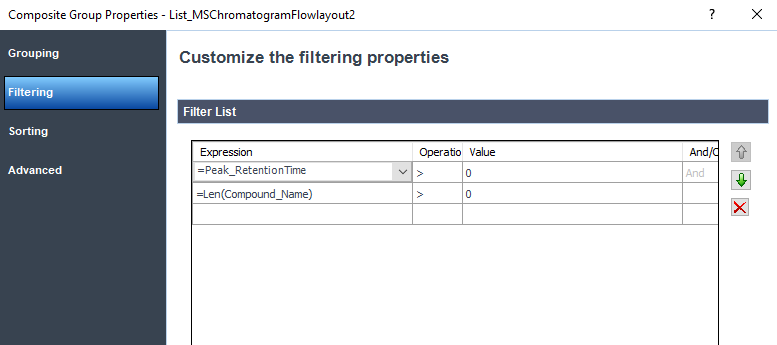
I erase the Len(compound_name) filter at the chromatogram and there is no sorting for this section.
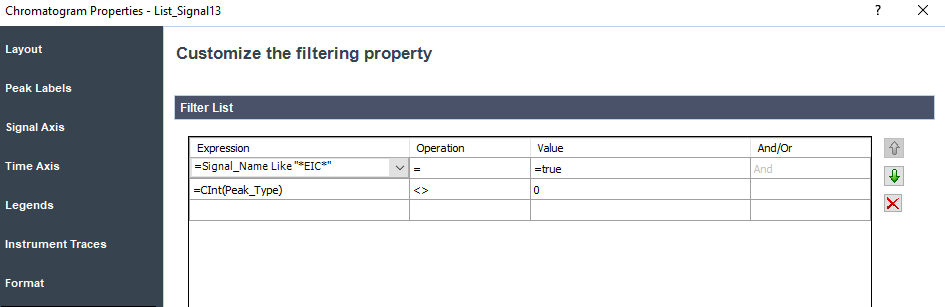
Here is the results:
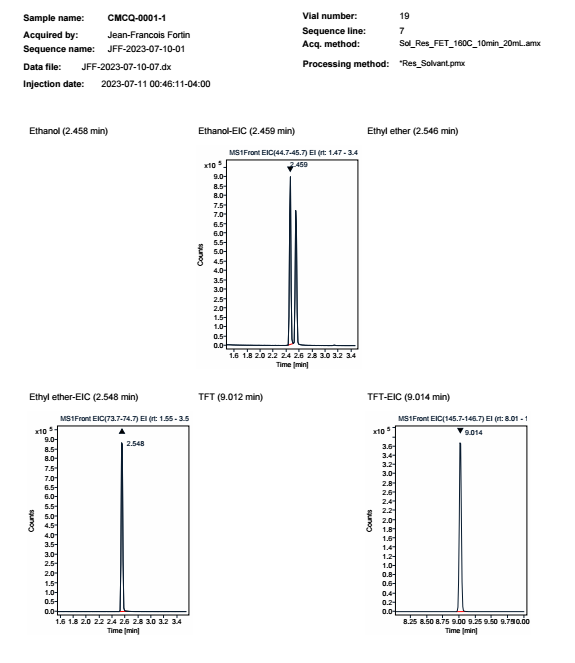
We can see that the filter EIC on the chromatogram doesn't work as final result as we can see blank chromato with no EIC as name compound.
I will try to add the EIC filter again on the outside filter.
it works! Even if I erase the EIC filter at the chromatogram portion. Thank you for your help. I will saved this composite group. If I add this composite group into a sequence summary report instead of single injection report, would you recommend to ass sorting based on injection ID?
I finnaly got what I have wanted. Thank you again. I tried to add the Multiplier used in the calibration Compound table used for a specific calculation with samples into my sequence injection summary:
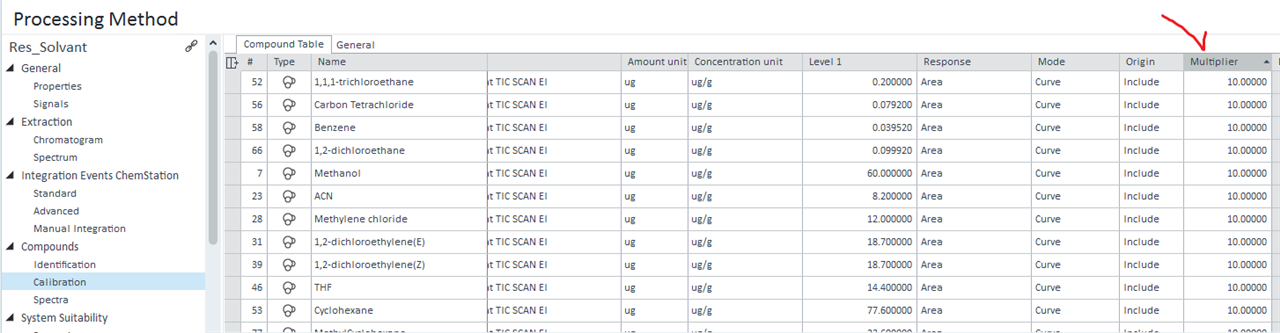
But I only found multipliers that included the 5 possible multipliers.
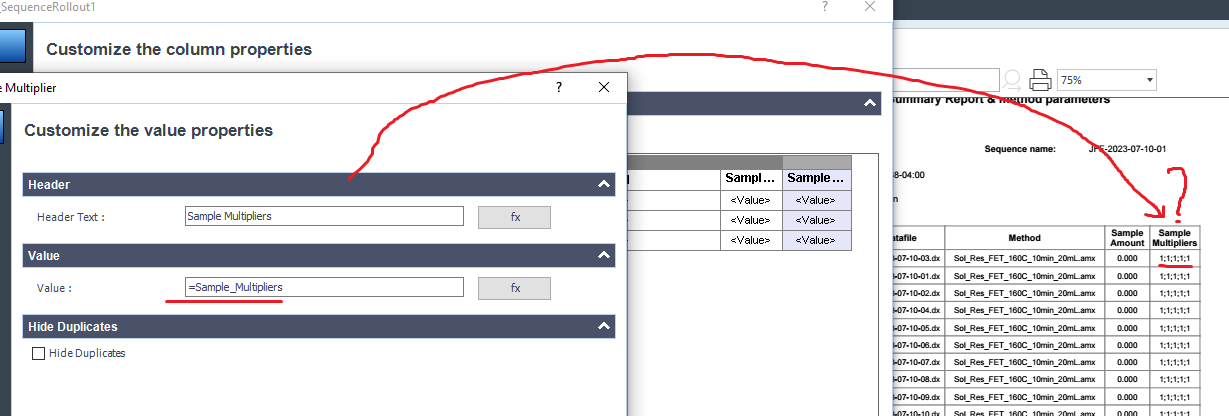
Do you know how to select a specific multiplier to add on the sequence summary?
Do you know how to change the response value of the courve into raw date ms counts?
Right now it's the 100% of response relatec to the 1 calibration point.
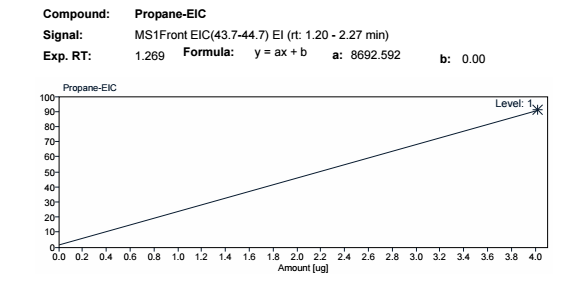
Do you know how to change the response value of the courve into raw date ms counts?
Right now it's the 100% of response relatec to the 1 calibration point.

If you don't want it to show only one calibration point, then you need to filter it to show only the last calibration injection. What I've done to resolve this so far is in a different field go filter by sample type for only cal standards, sorted by sample_OrderNo, then store the last sample_orderNo. Then in another field go filter by Sample_OrderNo, sort by injection_AcquisitionOrderNo, then store the last Injection_AcquisitionOrderNo. Then in the calibration curve filter by the stored Sample_OrderNo and Injection_AcquisitionOrderNo. That should give you the calibration curve as of the last calibration injection. If you want to see how it works better, just repeat on injection_ID and you will see the calibration curve build up one point at a time.
Also, I think your response is by percent because you have either the "Normalize to" or "Calculate mass %" boxes checked in the calibration general settings. I've included a snip to show where they are. I recommend playing around with those to see your options. I haven't done much with them so I can't help much more than that.
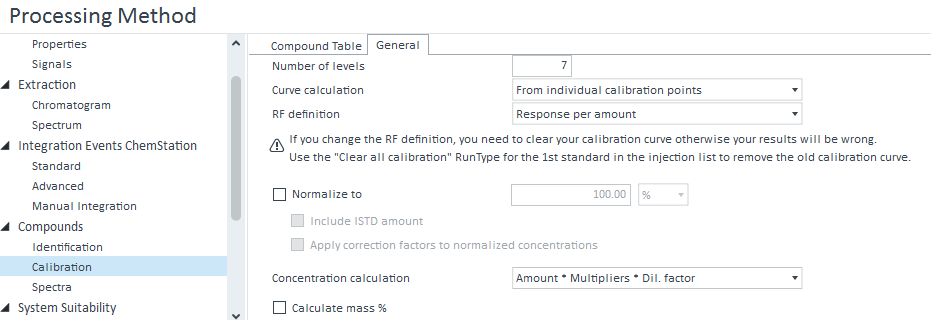
I need the % mass because the other option do not take into account the sample weight that I can put in my injection list for process.
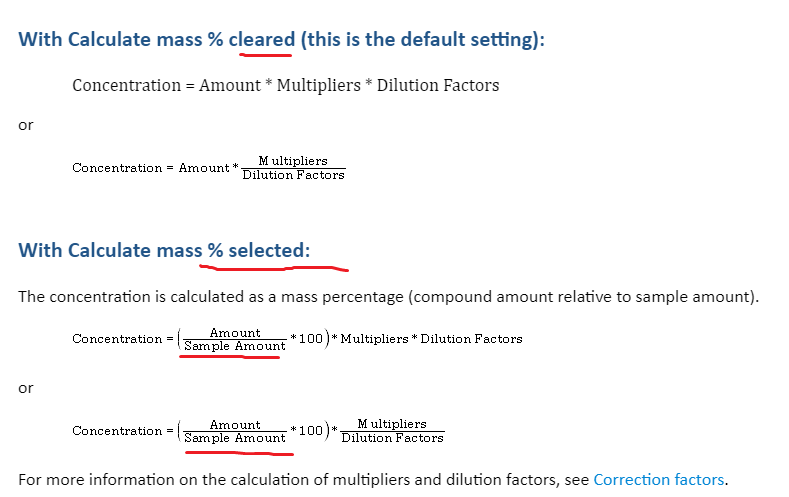
So the method will ask to the analyst to put sample weight in mg in the injection list. The concentration will be the amount founded by the one point curve (limit test) in ug divided by sample weight. I need to add multiplier to transform mg of sample into gram. The conversion scale is 1000 but since I already multiplied by 100, I have to add a 10 fold multiplier.
And it works perfectly. I can show the concentration calculated like that in ppm (ug solvant/g of sample)
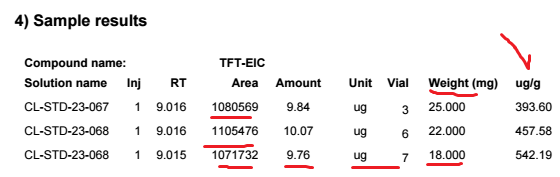
9.84 ug founded with my one point curve is then divided by the sample weight 25 mg gives 0.3936 ug/mg. In order to transform it into PPM (ug/g) we need to multipliy by 1000. 394 ppm and that's what we obtained. So the calculation is perfect without doing any custom calculation. Very simple.
But the problem is that I can't show my multiplier used in my sequence table into the sequence summary table in my report for our data revisor.
Also, by choosing this % mass, my response axis is in % amount based on calibration point. I can live with that if we can't change it to raw unit counts.
But I need to find a way to show my multiplier factor in my sequence table.
I'll do a little digging to see if I can find the field name. I've never had to reference this before so I'm unsure if you even can. I'll reply back if I find something