Hi,
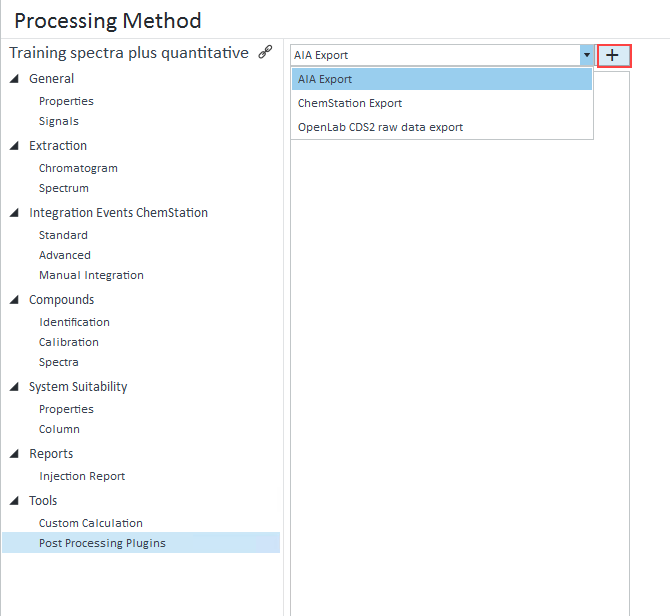
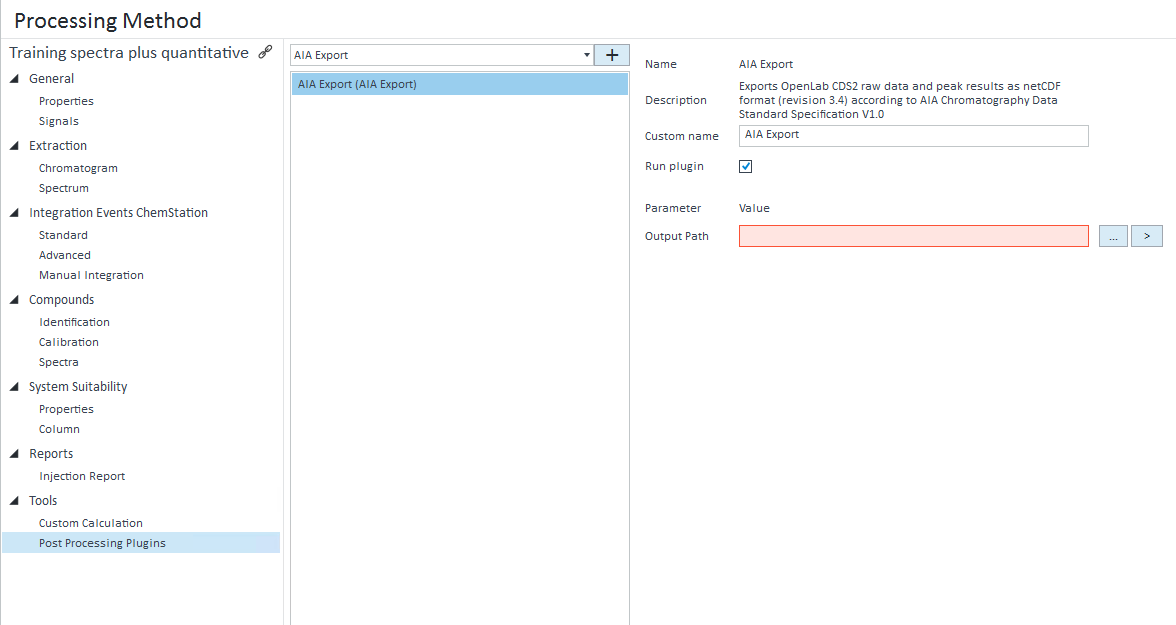
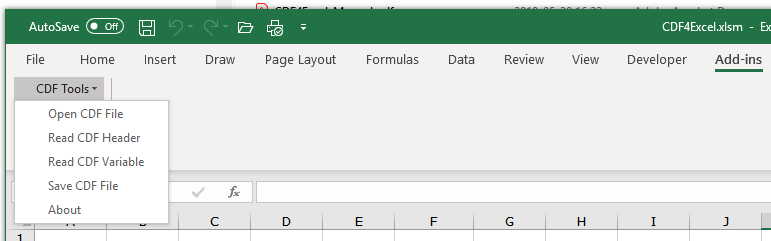
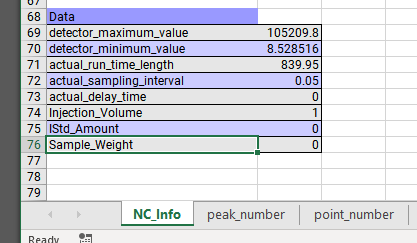
I'm looking to export the chromatogram as a CSV as chemstation was able to do. I want just the raw time vs. response. I don't want any peaks/areas/area% etc. Nothing that is a result of processing. OpenLabs manual suggest a method, but the only options are EMF and PNG. I don't want a graphical picture, I want the raw x,y values.
We use this exported data for performance verification through our own software and data analysis. Is this possible to achieve with the OpenLabs 2.2 CDS Workstation?
Thanks!