Greetings Agilent Community,
I see there are other posts that relate to this mysterious error, but none address my specific issue.
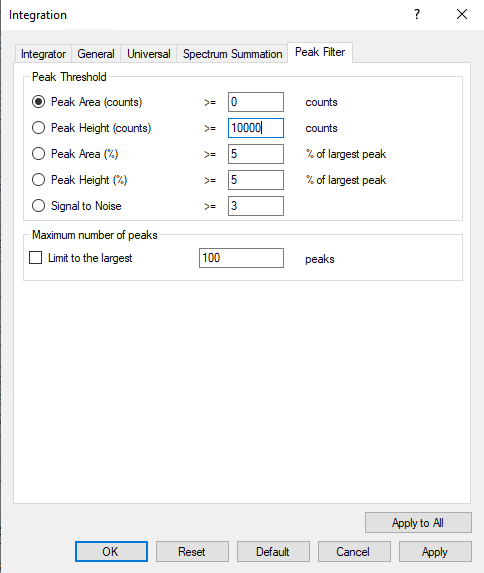
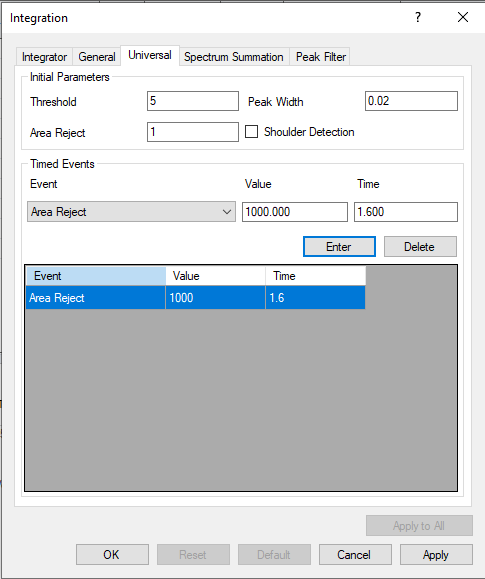
My task is simple: correct the default integration applied to blank samples either by zeroing the peak or manual integration. However, I am often thwarted while attempting to redraw the integration limits for my compounds. An error is thrown that reads "Manual integration failed for compound X in sample Y. Index was outside the bounds of the array." Alternatively, in other instances I do not receive this error, but the integration limits that I draw do not stick. Instead, one limit will snap to the very edge of the integration window and refuse to be positioned elsewhere. Very strange. Not sure if both phenomena have the same root cause, but they are happening concurrently. Has anyone else had this experience or know of a solution to my integration frustrations?
As always, your attention, insight, and advice are greatly appreciated!
Thank you,
Mike
MassHunter Quantitative Analysis for QQQ, Version 10.2, Build 10.2.733.8