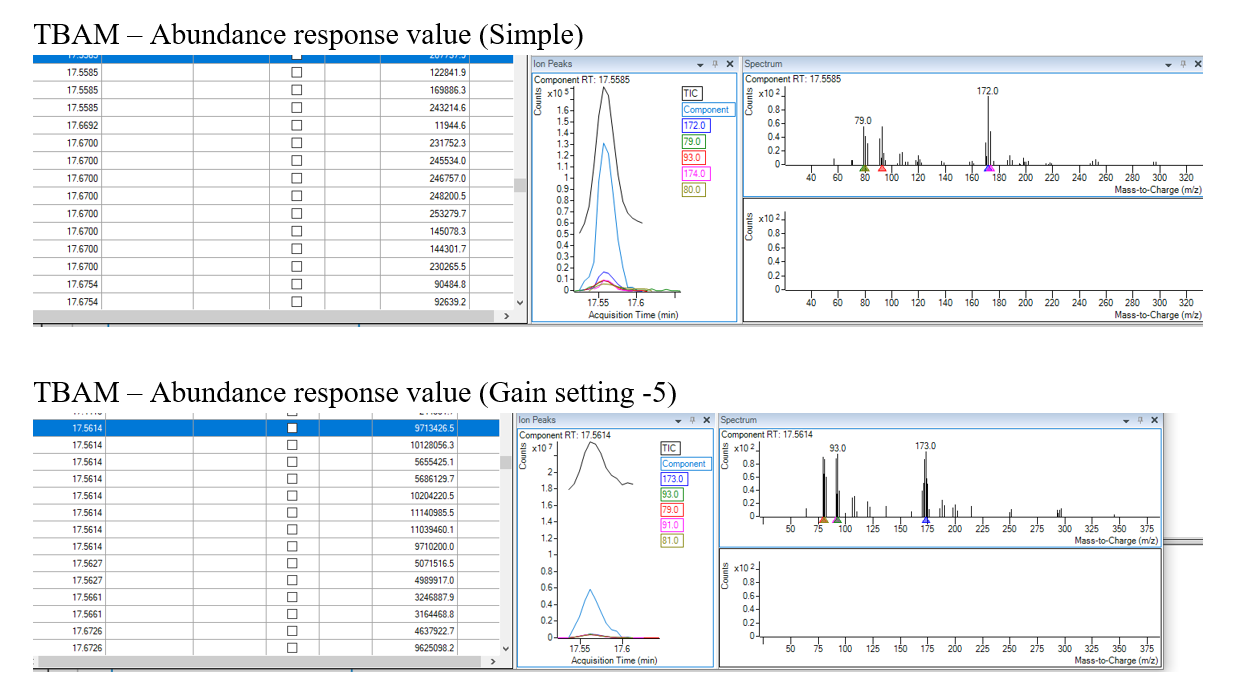
Hi,
I am developing an MRM (multiple reaction monitoring) method for Haloacetamides (HAMs). I've completed the following steps:
(1) Ran the full scan and identified the peaks using Unknown Analysis
(2) The HAMs I am working with are not present in the NIST library (except for one), so I made a custom library after completing the analysis.
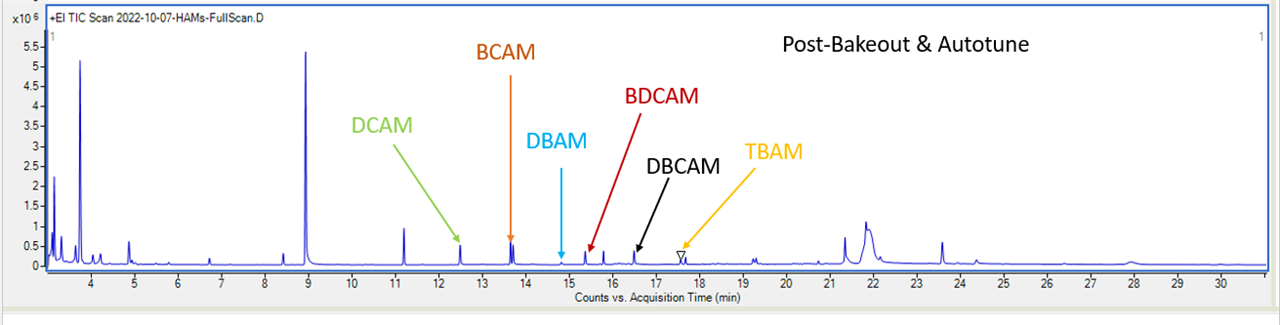
(3) In total, I have 6 HAMs - the peaks for them are pretty small, as can be seen in the chromatogram here:
(DCAM - Dichloroacetamide) (BCAM - Bromocholoracetamide) (DBAM - Dibromoacetmaide) (BDCAM - Bromodichloroacetamide) (DBCAM - Dibromochloroacetamide) (TBAM - Tribromoacetamide)
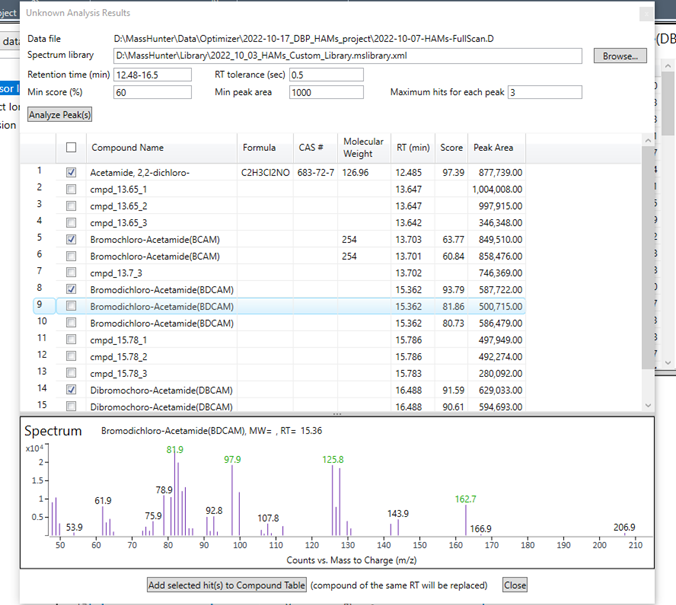
(4) Now, when I have made the library, and I am proceeding to the step of adding identifying the precursor ions in the Optimizer software, it is only picking up the 4 compounds (DCAM, BCAM, BDCAM, DBCAM) and not the TBAM and DBAM
Here is a screenshot from the Optimizer menu where I am using these settings to analyze peaks from a full-scan data file (the RT window that I am using is from 3 to 20)
And here is the library that I have made:
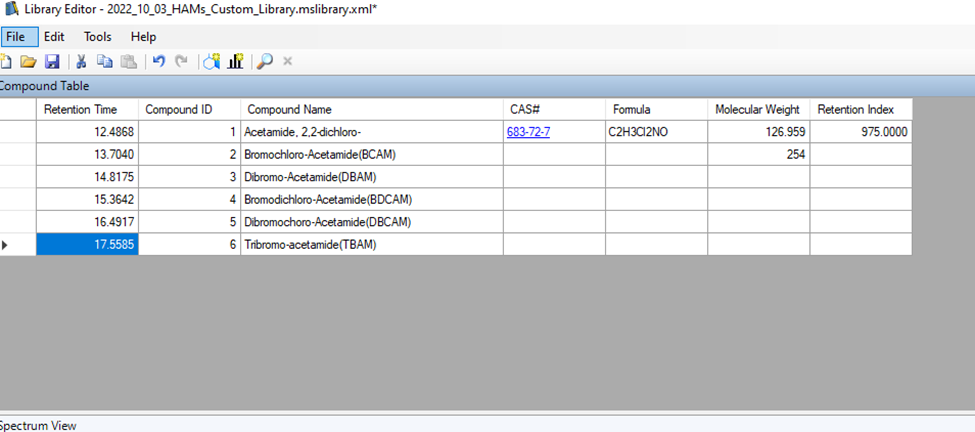
I am preparing the HAMs standards in 1mg/L concentration in methanol (1mL) and then diluting it with MTBE because my primary elution solvent is MTBE.
What settings can I change, try, or take steps to get the optimizer to pick these two compounds (TBAM and DBAM)?
Any help would be highly appreciated