Let me begin by saying I am totally new to using the calibration tab in OpenLab software. My lab currently uses a 1 point calibration curve and excel spreadsheets to manually process the data. I am in the process of validating a 5 point calibration curve and would like to use the software instead of excel to do the calculations.
I've watched the video on creating a calibration curve with an internal standard and understand the concept of the calibration table. However, the numbers I'm getting in the software do NOT match what I'm seeing on my excel spreadsheets and at times I'm unsure of where the numbers in OpenLab are even coming from.
I've tried looking for other videos/tutorials about how to proceed after a calibration table is established but can't find any information. Are there any videos/tutorials on the next steps after a calibration curve is established? I can set up the table, but then what? How do I use it?
I am using an internal standard and running 6 calibrators in triplicate (1 point will eventually be dropped) along with 3 controls.
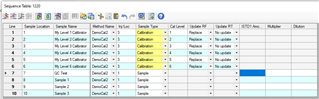
1. I need the software to ratio the std area and the IS area then average the ratios for the 3 runs. How should set up my calibration table? Should the first run be "replaced" or all 3 "averaged" for each level? Once the calibration table is established, how do I check that the software is averaging the ratios?
2. How do I save the calibration curve from a particular day's runs? How do I access it again w/o having to reapply all of the data files to the table?
3. How does batching work? How do I specify a particular calibration curve for the batch?
4. Do I need to specify in the sequence which samples are calibrators/controls? Where is this information used?
That's just the tip of the iceburg! I have SO many questions.....and that's before I even get to reporting options.