Hello
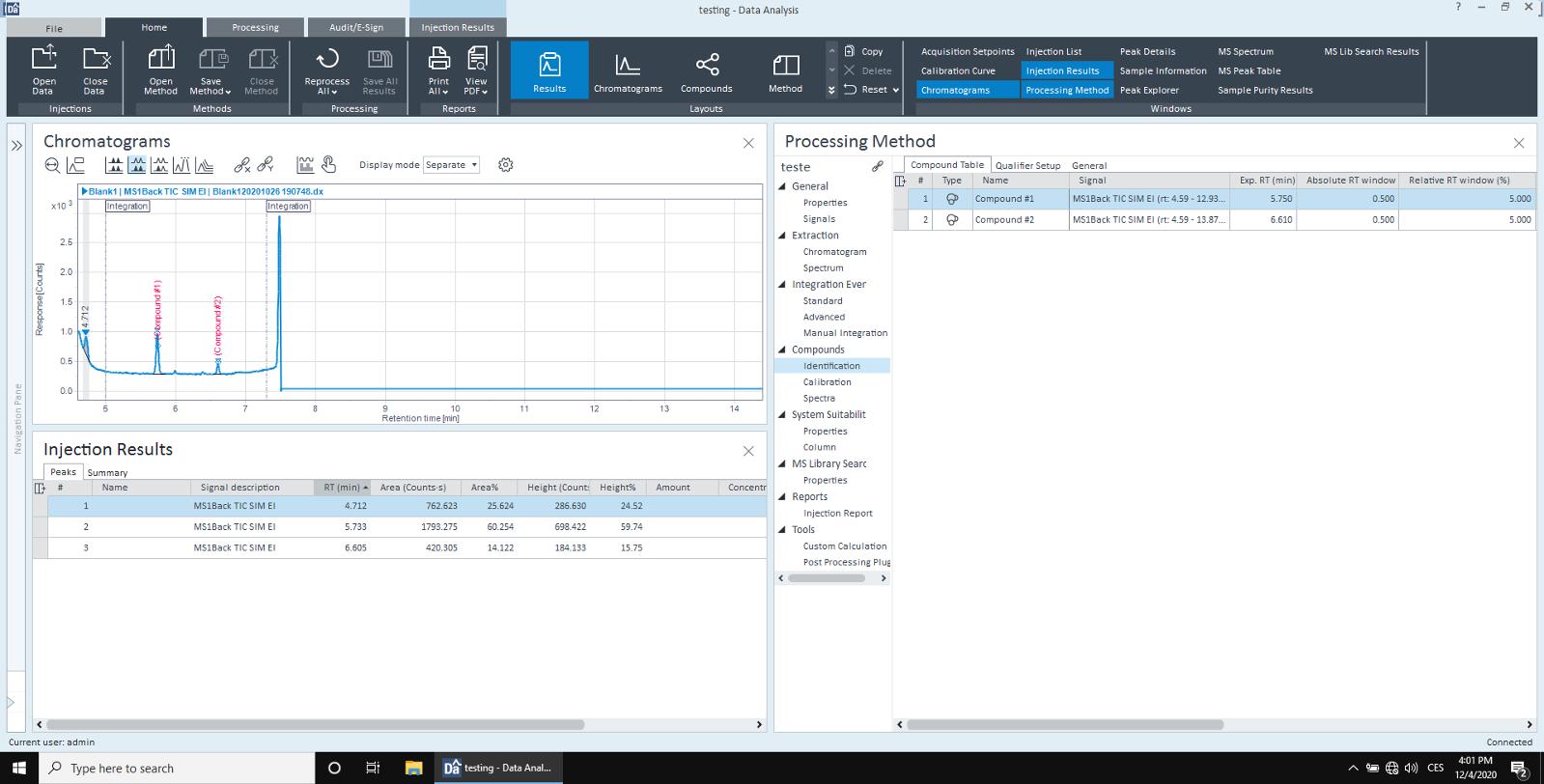
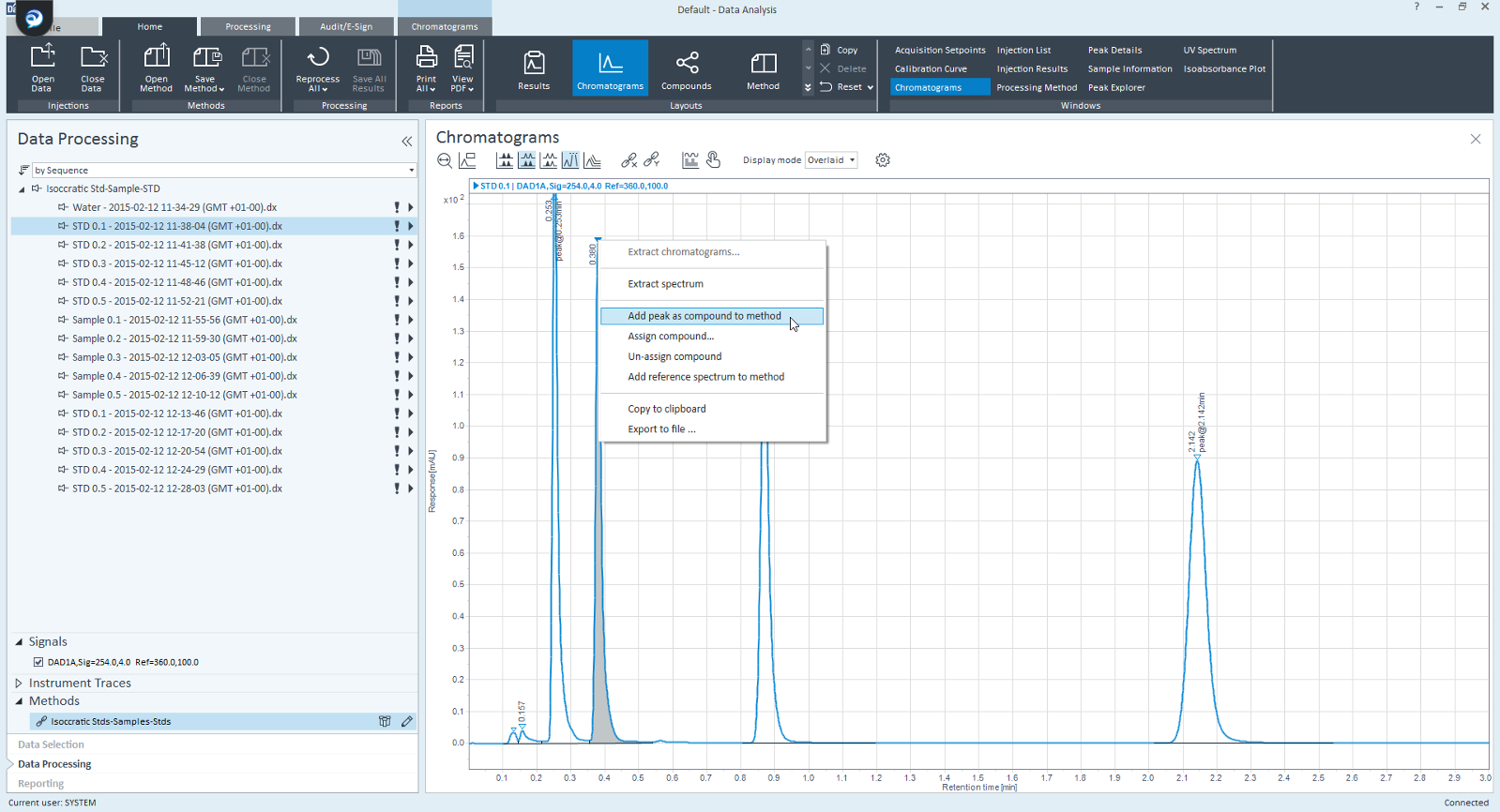
I have a SIM acquisition method and collected data. I need to create TIC compounds and calculate resolution, for whatever reason.
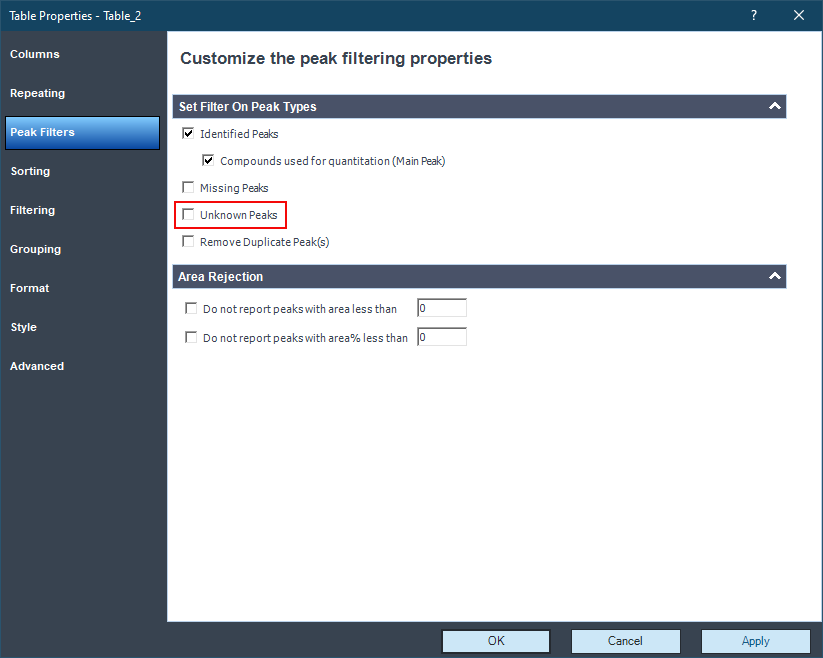
However, in the injection results, the NAME column is empty. I dont see these compounds in reports either.
thank you
openlab cds 2.5