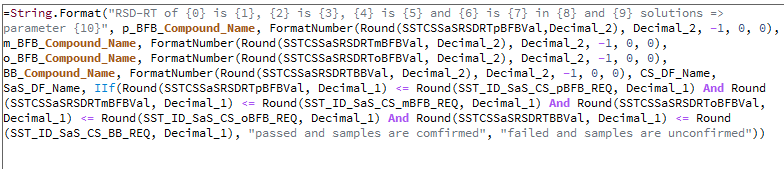
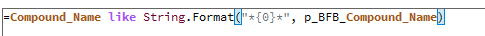Hello guys,
I met some kind of logic problem. Let me show you my situation: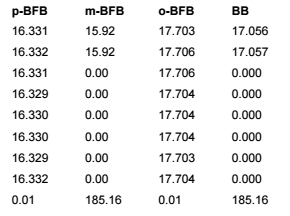
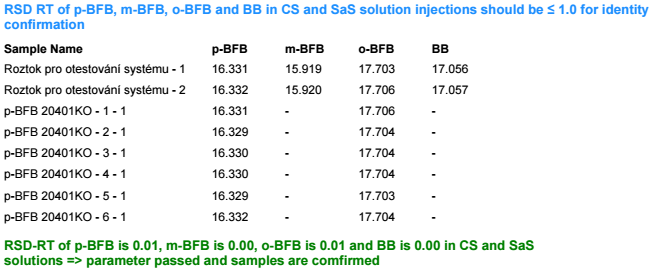
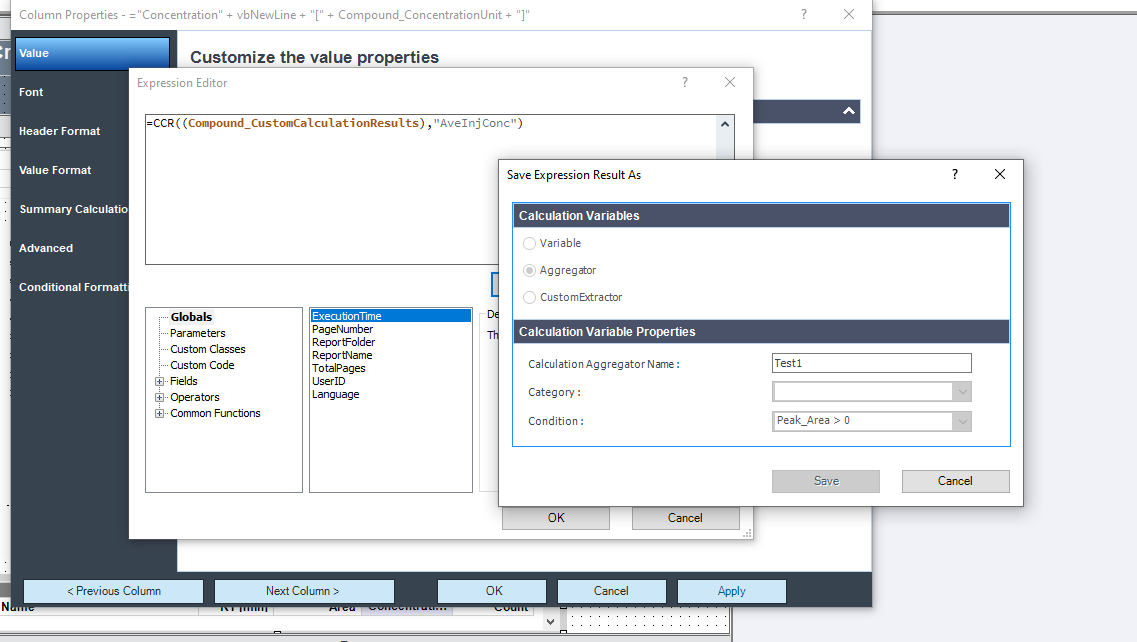
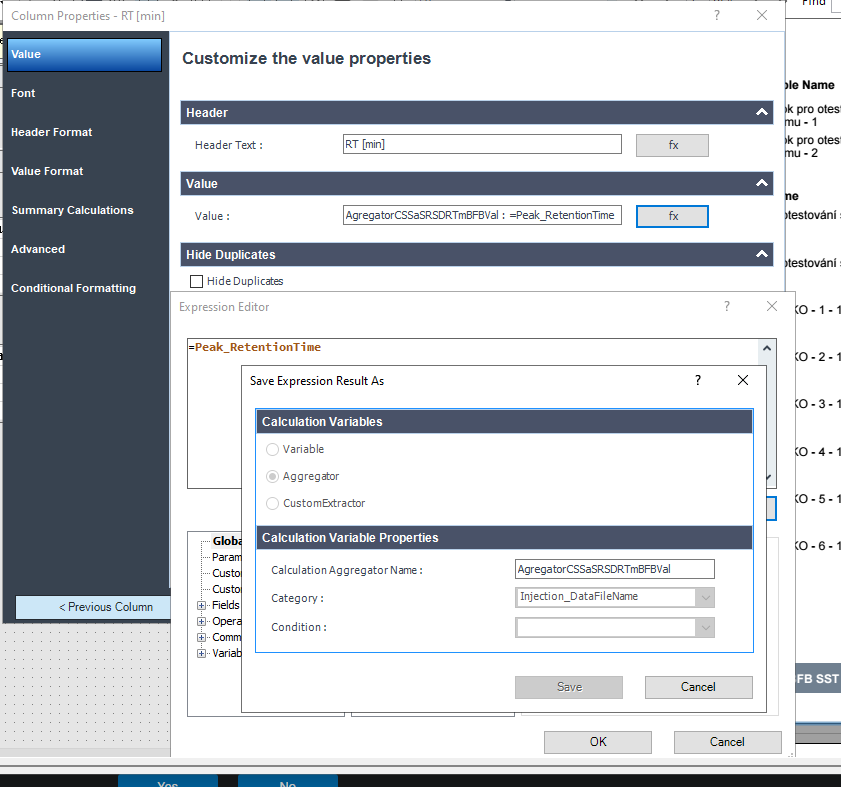
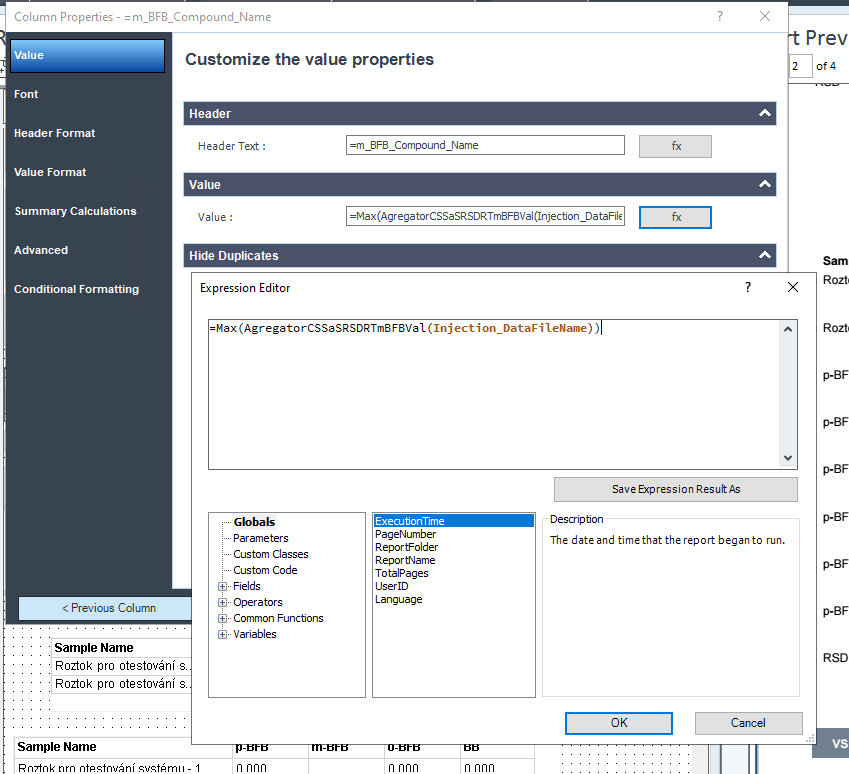
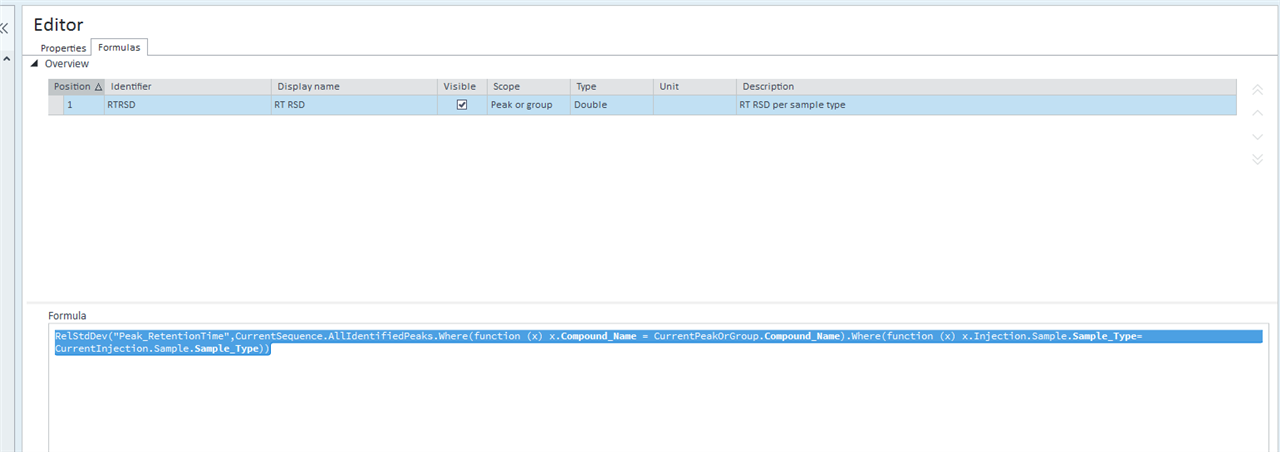
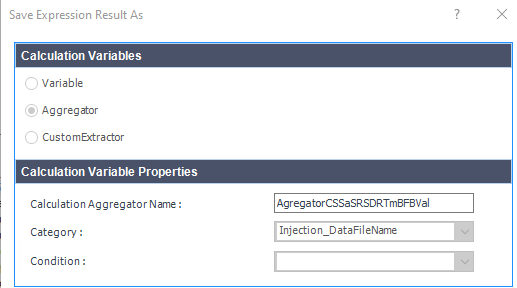
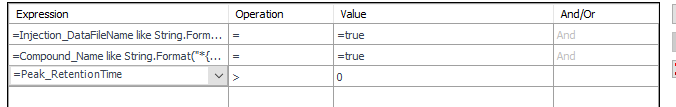
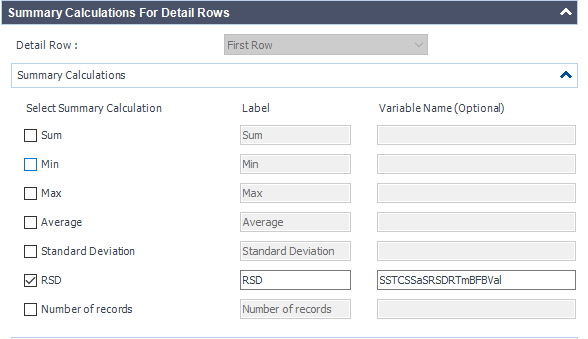
This table is simply a results table filtered for one main compound (p-BFB) and I added columns with other compounds by datafile aggregator. The problem is when I am comparing some solutions, where compound m-BFB or BB is missing. I expected lines to be empty objects. Instead, OpenLab is converting empty values into 0 so if I want to calculate RSD from all occurrences it is calculating with 0. In my example, I am calculating the RSD of RT and when I have 2 standard injections and some sample injections in the set I have lines defined by the main peak and I am also going to have at least 2 impurities RT value but sometimes is sample clean and I do not have any peaks so I get 0.
Does somebody know some trick how to handle this situation?
Thank you guys