Greetings forum denizens,
I'm running on Openlab CDS Version C.01.10[201] . I'm (still) working on setting up our GC for pesticide analysis. We're looking for about 30 compounds. As many of you know, these can elute very closely. To make things easier, we are using two sets of standards: one compound mixture is "A', the other is "B". I have made calibration standard for both and entered them. To reduce paperwork, my boss would like to replicate an old method we used twenty years on a 5890 (!) using an old, old version of Chemstation. This method used one calibration that combined those of standards "A" and "B" ; one curve with one report format that worked for both standards.
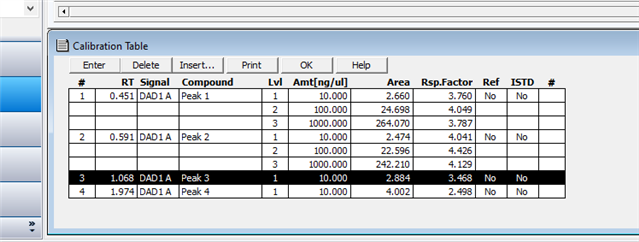
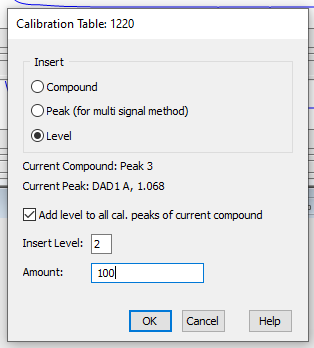
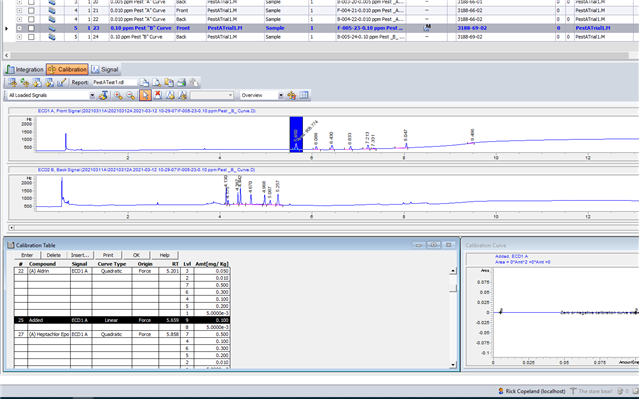
My question is simple: How can I combine the curves for both "A" and "B" in a single pesticide calibration? I played around and was unable to do so. I tried adding peaks. I tried adding one standard as separate levels (a dumb idea, but figured it was worth a try). I even hit upon the idea of overlaying chromatograms and trying to calibrate off this combination. Believe it or not, it worked for the first cal level, but I couldn't add more levels to the curve. Whether I can do this isn't going to bring calamity on my GC world. If worse comes to worse I can just use separate curves for the "A" and "B" standards. But we were capable of doing this on Chemstation , so I would think I can do it with my current OpenlLab CDS.