Hello. I wanted to see if anyone has come across or written a macro to export a batch of files into csv natively within Openlab C.01.10? There was a macro created for this purpose in older versions of ChemStation that was very useful.
Hello. I wanted to see if anyone has come across or written a macro to export a batch of files into csv natively within Openlab C.01.10? There was a macro created for this purpose in older versions of ChemStation that was very useful.
Hello,
Would you mind clarifying what you mean by exporting to csv? Are you looking for a summary report for a batch of samples in a .csv format or do you want the raw data x-y coordinates from the detector signal exported for each injection in a batch to a .csv file?
I am looking for the raw data x-y coordinates for each injection in the batch following integration in order to create an integration table with areas, percentages, and retention times as well as overlays of each injection via macros that I have made in excel
Thanks a lot, howard_sanford for the prompt reply. It works now and initially, the macro file was somehow named as .mac.txt file rather than .mac file. Now it generates a list of separate cvs files with each sample having three signal files (DAD1A, DAD1B, FLD1A). Just wondering if there is a way of only generating one signal file like DAD1A. I greatly appreciate it.
Chao
The macro prints all loaded chromatograms, so you can edit your method to only load the DAD1A chromatogram and that will be the only one exported. Otherwise you could edit the macro so that a particular chromatogram type is the only one exported.
Hi howard_sanford,
first of all thank you for posting this macro, it has been very helpful for me and my colleagues.
We were wondering if this macro could be implemented to obtain a single CSV file containing all the signals from each sample in the run, so that we don't have to open each individual file and merge the single signals for every sample (we work with 50 samples per run and it is quite painful to process them). Ideally it would be great to have also a single file containing all the RTs and peak areas per sample, again to speed up the processing and get the concentration of the compounds.
This macro would save us an incredible amount of time if it would be possible to obtain.
Thank you very much in advance!
Hello mfp_sadler ,
I'm glad you found that useful. The previous change I made was a very minor change to a very old user contributed file. I don't believe this macro could be easily modified to put all the results in one file. You could check the macro given as the other solution to see if that may be easier to modify. Or it is possible another user has something that they may be able to share that is closer to what you want.
You could also try to search for a batch file or shell script that could concatenate the files and do something like placing the file name in between each section.
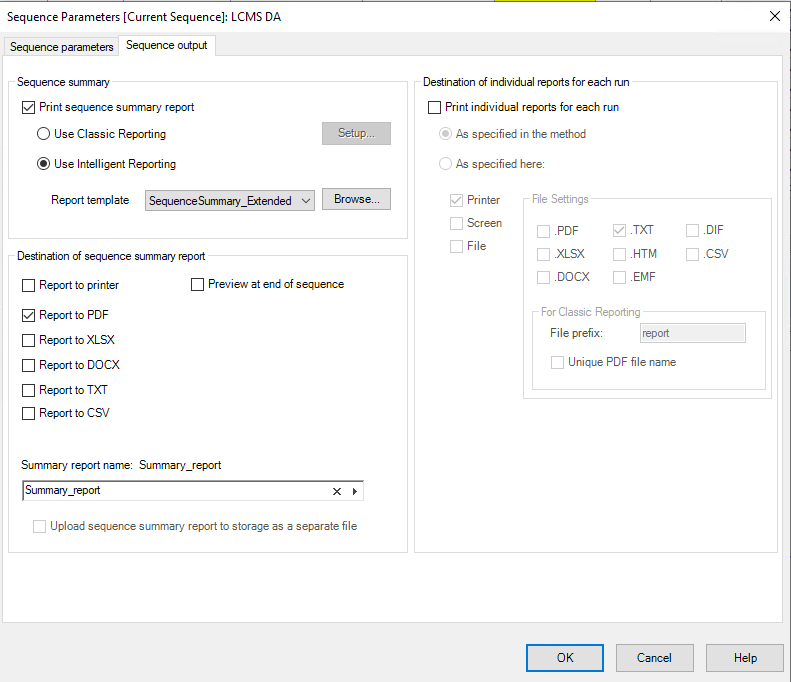
For getting all the RTs and peak areas into one file you could investigate using a Sequence Summary report. This can be set in the Sequence Parameters under Sequence Output.
Hi Howard,
This macro is amazing, but I'm wondering if you have something exactly like this (CSVxport.mac) that exports the X-axis (time values) and Y-axis together in one shoot?
Thanks!
Hello Lucas_S ,
Based on your follow-up explanation in this discussion -
I don't believe this macro could be easily modified to do something like that, with the x values in one column and each run's y values in subsequent columns. This was a user contributed macro that I made only slight modifications to and I'm not aware of any similar ones other than the other one mentioned in this discussion.
Hello Lucas_S ,
Based on your follow-up explanation in this discussion -
I don't believe this macro could be easily modified to do something like that, with the x values in one column and each run's y values in subsequent columns. This was a user contributed macro that I made only slight modifications to and I'm not aware of any similar ones other than the other one mentioned in this discussion.