Hi,
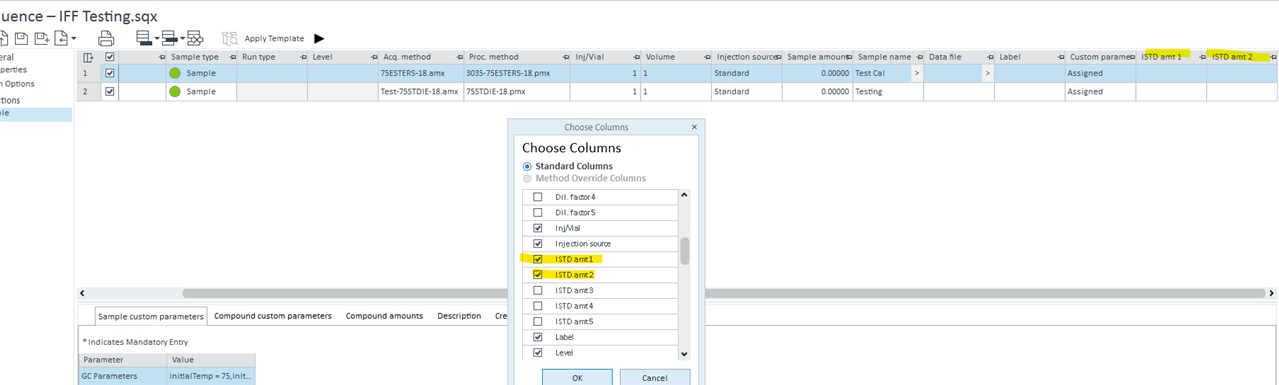
I am setting a method in OpenLab CDS where the ISTD (calbration) amount is different for each run. I am wondering if it is possible if we key in the ISTD weight in the sequence table, and the amount could be auto-calculated when I review the chromatogram?
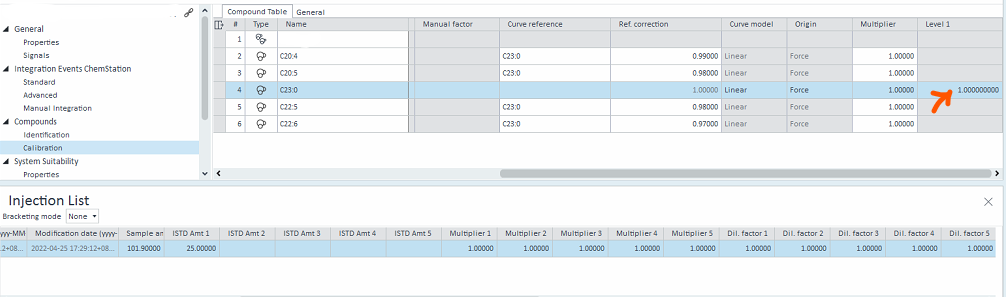
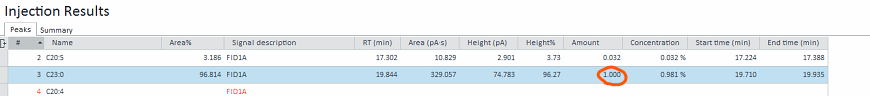
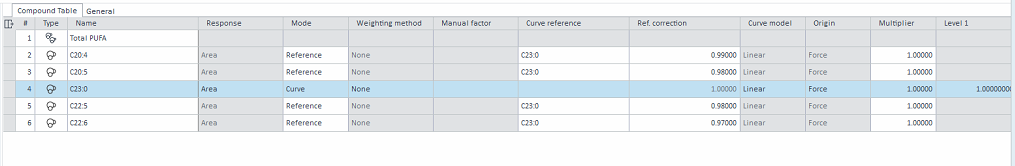
C23:0 is the ISTD and would be spiked into each sample, hence the amount spiked in would be different for each run. The amount of C23:0 = ISTD amount / 25
Thus my question is, is it possible to let the software auto calculate the amount of C23:0 (which will change the Level amount automatically for each run), by entering the ISTD amount and multiplier?