Hello!
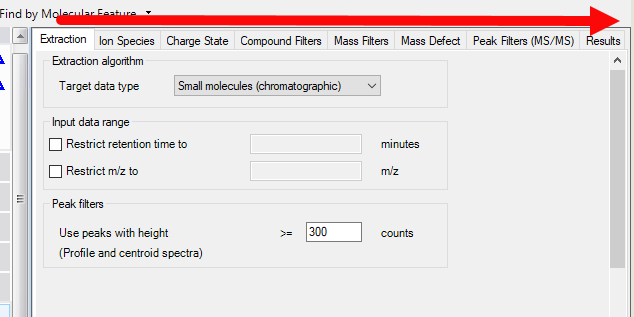
I want to import spectra in my PCD (just formula and mass included). I first ran my compound in targeted MSMS mode. Then I performed a Qual analysis (workflow: compound discovery, find by targeted MSMS). I exported the spectra as .cef file and then I added them to my PCD. The spectra don't look good though. There is a lot of noise and unknowns. Then I tried to run a find by molecular feature in Qual so that I can get the peak annotations but it seems that it identifies more than a thousand of compounds! In general, I am wondering what is the exact procedure if someone wants to add his own spectra in a PCD. Is there any technical note or something that describes in very detail the workflow to be followed.
Thank you in advance
Maria