Hi,
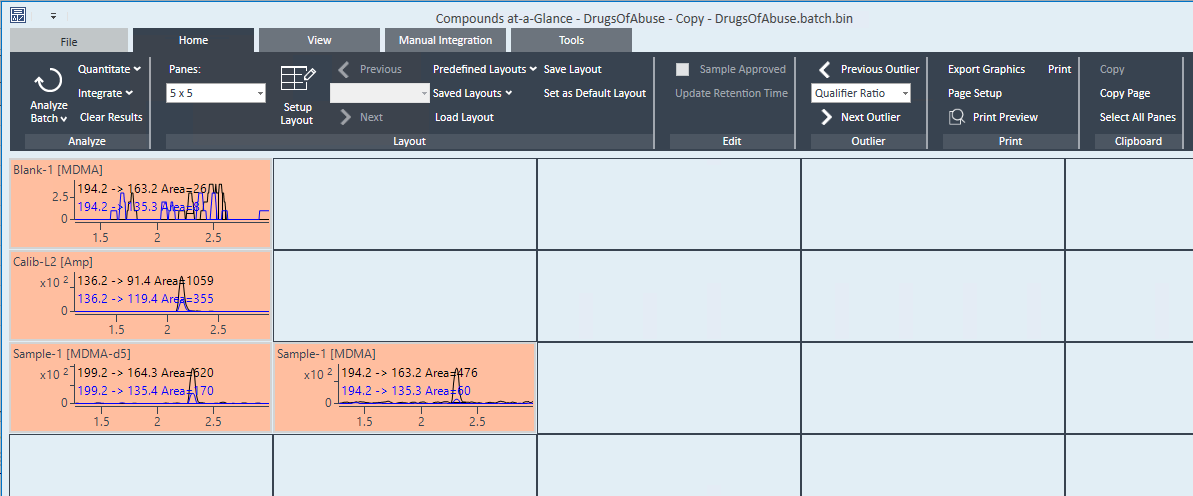
I am using the MassHunter Quantitative Analysis 10 software to quantify GC-MS data from plant samples, and I have standards and standard curves which I am using to quantify individual compounds.
Some of my standards are not expected to be present in some samples, but they may have peaks at about the same retention time as other compounds. In the case of limonene and beta-phellandrene, they cannot be separated by retention time, but the qualifier ion ratios are different so it is easy to distinguish them.
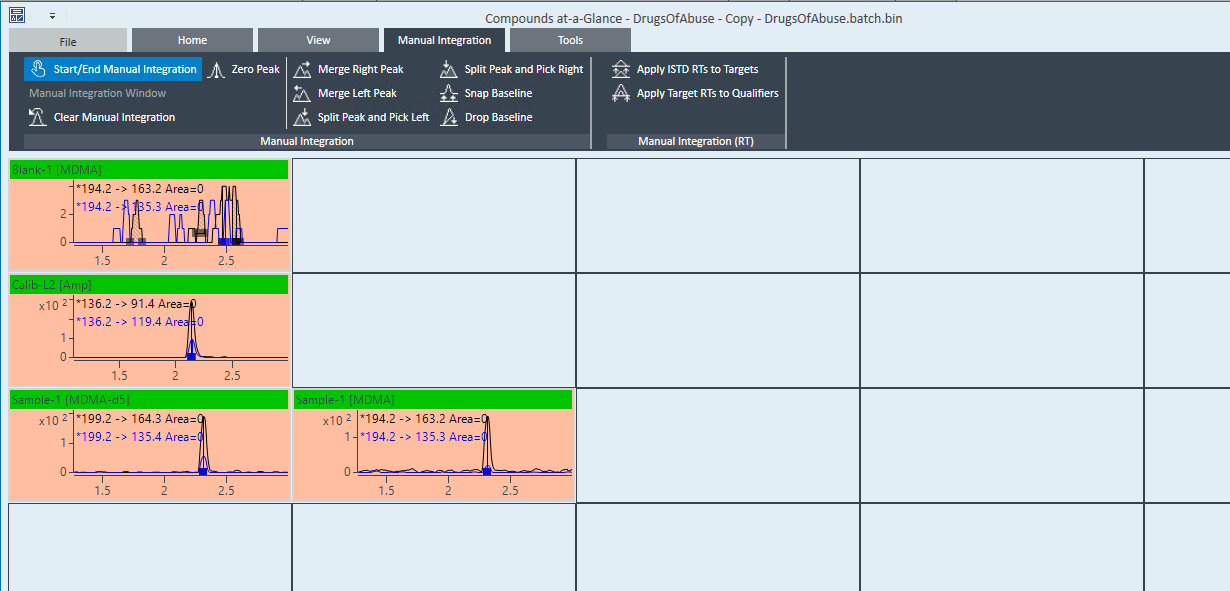
The problem that I am having is that for a sample which should ONLY have limonene, the software identifies the single limonene peak as both limonene AND beta-phellandrene, even though the qualifier ion ratios are completely out of bounds for beta-phellandrene and they are perfect for limonene.
I have already set all compounds in the method to use "Close RT with qualifiers". I've set up the qualifier ion ratios to have strict uncertainty percentages. The ratios just get highlighted in the results but the peaks are still being identified as the wrong compound. Is there anything else that I can do to prevent misidentification? I don't see a way to specify "minimum number of qualifiers" or anything like that, and I'm getting false-positives for other compounds because of this as well. Even in samples where there are no matching qualifier ions, the software still gives a target match and a concentration.
What have I set up wrong here? Have I overlooked something?
Thanks!