Hello all.
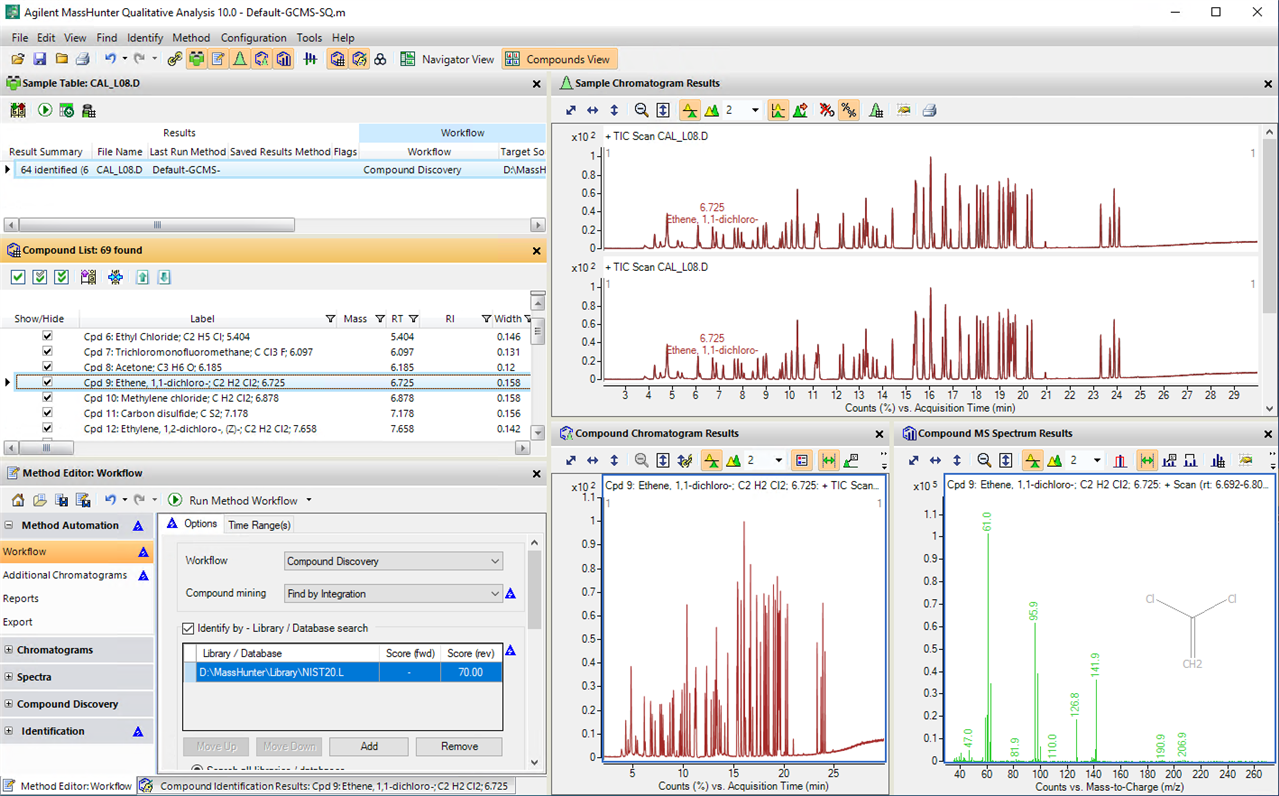
We just got a new GC-MS and it uses the Masshunter software for the processing the results.
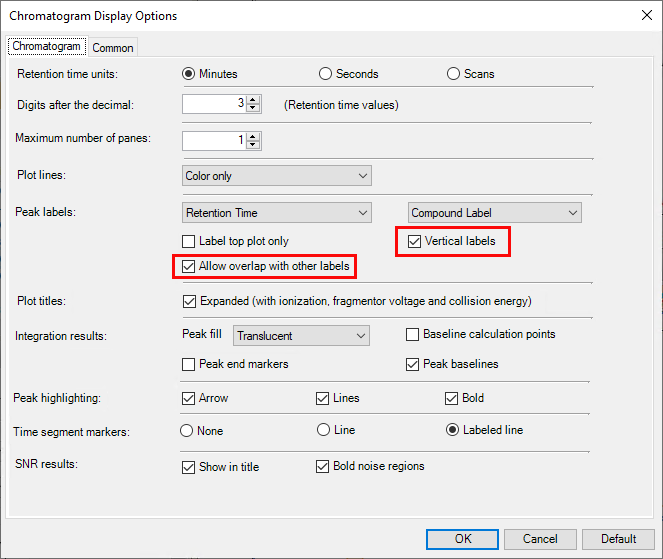
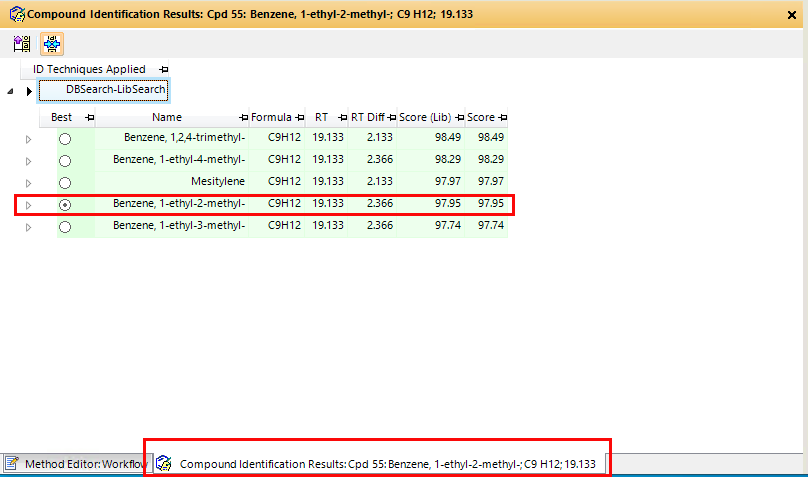
We have an analysis run of 27 compounds, so when I'm processing the spectra in Masshunter Qualitative, it's hard to always remember which peak is in that exact RT, specially when you have more than one peak close together. So I wanted to add labels to all peaks according to RT. I was able to find how to manually add comments/labels but that would mean that I had to do it for each and all samples processed. Is there a way to program it to automatically add the labels to all spectra that I open with the software ?
For the quantitative analysis:
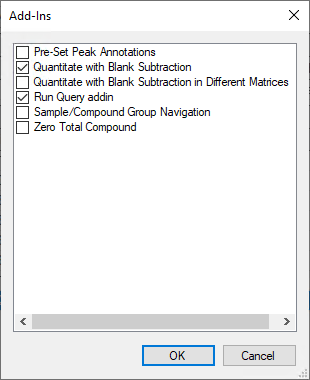
I tried a test run consisting of only a blank, a QC run as QC, and the same vial run as sample. When processing the results I saw that, although i had responses in the blank, the software was not subtracting that amount from the samples when giving the final concentration. How can I set it up so it does? I tried to use the custom calculation column, but could not find a way to define the calculation steps.
I appreciate any help.