Hi all
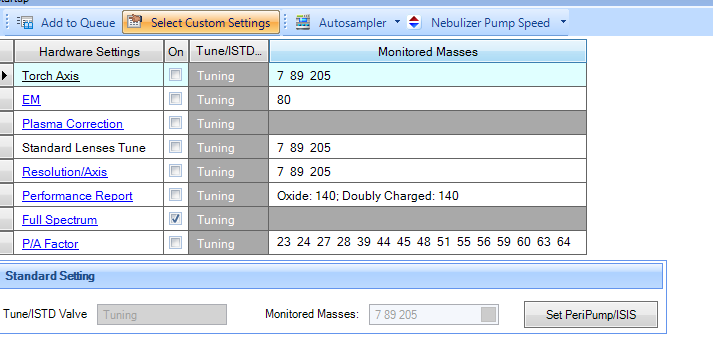
I have run a full spectrum on startup to check for contaminations. I was surprised with the high counts measured for most elements.
Does that look right? Am I missing something? The first line is the full spectrum and below is the calibration run after it.